In ad dition, selected of those transcriptionally opposing histone modifications have been proven to become mutually exclusive at identical websites inside the promoter regions of energetic vs. repressed alleles at imprinted loci suggesting a poten tially robust technique for looking for candidate imprinted loci, independent of expression based analyses determined by DNA sequence variation, this kind of as single nucleotide poly morphisms, Preceding searches for imprinted genes in marsupials have focused on a small number of loci which might be previously identified to be imprinted in eutherians, and only a few of these have sought to describe DNA methylation and histone modification profiles of CpG islands at putative promoter regions, Clear evidence of dif ferential DNA methylation at marsupial imprinted genes has become observed at only two loci, and only the DMR with the IGF2 H19 imprinting cluster continues to be proven to manage transcription of the marsupial imprinted gene, In addition, the marsupial X chromosome, which exhibits pa ternal imprinting for most loci in females, is strongly defi cient in CpG island methylation, Information addressing histone modifications at promoters and CpG islands of marsupial imprinted genes are incredibly constrained, with only two histone modifications, H3K3me2 and H3K9me3, examined for opossum Igf2r, Htr2A, and L3mbtl.
These genes exhibit enrichment of H3K4me2 but not H3K9me3 at promoters, Chromosome degree immunofluorescence analyses of wallaby, opossum, and brush tailed possum X chromosomes, utilizing antibodies to particular histone modifications, have proven correlations among a number of repressive and activating histone marks for the inactive and active X chromosomes, respectively, constant with a possible purpose for selleck chemical BIX01294 histone modification states while in the transcriptional regulation of genes on the marsupial supplier Mocetinostat X, However, these cytologic approaches lack electrical power to resolve the locations of modified histones on scales a great deal below the chromosome band degree, so are unable to determine correlations between histone modification distributions and expression states of indi vidual genes.
Taking advantage of constantly enhancing subsequent gene ration sequencing technologies plus the high quality draft assembly with the M. domestica genome,  we’re now capable to hunt for marsupial exact imprinted genes and analyze fundamental signals of imprinting on the genome broad basis. To complete this, we conducted reciprocal crosses of animals from two M.
we’re now capable to hunt for marsupial exact imprinted genes and analyze fundamental signals of imprinting on the genome broad basis. To complete this, we conducted reciprocal crosses of animals from two M.
Monthly Archives: May 2014
23 The examination was performed applying the siggenes package d
23. The analysis was performed working with the siggenes bundle via the R application surroundings for sta tistical computing and graphics, Transcripts exhibiting appreciably up regulated expression have been annotated applying Gene Ontology terms and hierarchical structure.The Blast2GO plan, which assigns the GO terms primarily based around the BLAST definitions, was utilized with an E worth 10 five degree. Northern blot analyses Northern blots were obtained using complete RNA extracted from T. harzianum CECT 2413 freeze dried mycelia col lected as described above. RNA separation, blot ting and hybridization had been carried out employing standard methods. Distinct DNA probes of every gene had been professional duced by PCR in the corresponding cDNA clone from our Trichoderma cDNA clone assortment, utilizing the primers T3 and T7 through the Uni Zap XR vector, The DNA probes have been P32 labelled implementing Ready to go DNA labelling beads and radioactive signals have been visualized using a PhosphorIm ager System, utilizing Quanti tyOne computer software.
Burkholderia pseudomallei, causal agent from the poten tially fatal condition melioidosis, is usually a metabolically versatile soil organism purchase GSK2118436 that has been classified as being a Category B biological threat through the CDC, Somewhat small is identified about its pathogenesis, virulence factors, the extent of diversity in natural populations, and host response. B. pseudomallei genome plasticity is connected with genomic island variation. The genome of B. pseudomallei K96243, as an example, features 16 genomic islands, at least 3 of which appear to get prophages, Its unclear whether all of these putative prophages are lively, whilst one was proven to be a productive bacteriophage. B.
pseudomallei isolates are genetically pretty various, and this het erogeneity may perhaps be due at the least in element to the tremendously vari in a position distribution of bacteriophages among strains, This kind of differences could provide certain strains survival strengths from the atmosphere along with the host, at the same time as clarify the variable U-95666E clinical presentation of melioidosis. Also raising concern like a possible biological weapon is definitely the very closely associated B. mallei, causal agent of the pri marily equine disease often called glanders, In contrast to B. pseudomallei, B. mallei is really a extremely specialized pathogen, not observed outside of the mammalian host in nat ure. B. mallei is often a host adapted clone of B. pseudomallei, and each of the B. mallei genome is just about identical to a set of genes within B. pseudomallei core genome. Nevertheless, moreover to its core genome B. pseudomallei is made up of various contiguous gene clusters that were deleted from B. mallei while in its evolution, B. thailandensis is one more closely relevant organism usually located during the same environmental samples as B.
Cells have been examined working with a fluorescence microscope a
Cells have been examined making use of a fluorescence microscope and all cells in a specified spot from the middle in the membrane have been counted. Scratch assay Cells have been seeded at a density of 450,000 cells per effectively in twelve very well cell culture plates. Soon after incubation for 24 hrs, the confluent cell monolayer was scraped using a pipette tip creating a scratch in every single well. Medium containing serum supplemented with TPA or inhibitors was added and cells had been incubated at 37 C. For experiments with siRNA, 70,000 cells were seeded in twelve nicely cell culture plates and handled with siRNA as described and 18 hrs following the final transfection, cell monolayers had been scratched. Cells have been photographed at unique time points along with the scratch area was measured utilizing ImageJ. Western blot 1. 0 ? 106 cells had been seeded in 60 mm cell culture dishes and incubated for 24 hours. Cells have been pre incubated for 1 h in serum free medium prior to stimulation.
Cells had been washed twice in PBS and lysed in RIPA buffer containing 401 ml protease inhibitors. Cells transfected with siRNA had been lysed during the exact same way 18 h following the last trans fection. Lysates had been centrifuged for 10 min at 14,000 ? g at 4 C. Proteins have been electrophoretically separated on the 10% NuPAGE Novex Bis Tris gel buy Cilengitide and trans ferred to a polyvinylidene diflouride membrane. For detection, membranes have been incubated with key antibodies towards phospho signaling transduction MARCKS. phospho Erk. Erk. MARCKS. PKC. PKC II. PKCor PKCfollowed by incubation by using a horserad ish peroxidase labelled secondary antibody. Horseradish peroxidase was thereafter visualised employing the SuperSignal technique as substrate. The chemoluminescence was detected having a CCD camera. Calculations and statistics IC50 values had been calculated by carrying out a curve match examination to your equation y A the place A will be the maximal impact and B may be the IC50 value.
Statistical 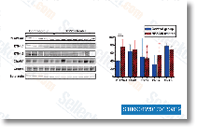 analyses have been performed by performing ANOVA followed by Duncans many array test utilizing p 0. 05 as degree of for significance. Success Activation of PKC stimulates migration of neuroblastoma cells To investigate a putative position of PKC in neuroblastoma cell motility, the migration of SK N BE C neuroblast oma cells was studied making use of transwell and scratch assays. SK N BE C cells have been seeded inside the upper wells in the transwell assay and have been permitted to migrate in direction of serum cost-free medium supplemented with sixteen nM from the PKC activator TPA. It is a treatment method that isn’t going to lead to morphological changes on the cells This demonstrated that TPA contributes to a doubling in the quantity of migrated cells. Because TPA can influence other proteins than PKC isoforms. PKC inhibitors had been incorporated with TPA in the reduce chamber to investigate if PKC activ ity mediates the TPA effect. Each the standard PKC inhibitor GF109203X plus the inhibitor from the classi cal isoforms, G6976, markedly decreased the TPA induced migration.
analyses have been performed by performing ANOVA followed by Duncans many array test utilizing p 0. 05 as degree of for significance. Success Activation of PKC stimulates migration of neuroblastoma cells To investigate a putative position of PKC in neuroblastoma cell motility, the migration of SK N BE C neuroblast oma cells was studied making use of transwell and scratch assays. SK N BE C cells have been seeded inside the upper wells in the transwell assay and have been permitted to migrate in direction of serum cost-free medium supplemented with sixteen nM from the PKC activator TPA. It is a treatment method that isn’t going to lead to morphological changes on the cells This demonstrated that TPA contributes to a doubling in the quantity of migrated cells. Because TPA can influence other proteins than PKC isoforms. PKC inhibitors had been incorporated with TPA in the reduce chamber to investigate if PKC activ ity mediates the TPA effect. Each the standard PKC inhibitor GF109203X plus the inhibitor from the classi cal isoforms, G6976, markedly decreased the TPA induced migration.
Then, 46 five mg of this solution, FSH33 PEG, was added to ten m
Then, 46. five mg of this product, FSH33 PEG, was added to 10 mL of two mg mL PEI remedy and magnetically stirred for 24 h at room temperature. The product, FSH33 PEG PEI, was dissolved and added in to the similar volume of plasmid DNA remedy drop by drop with magnetically stirring. The molar ratio of nitrogen from PEI to phosphate from pDNA was 25. The final complex, FSH33 PEG PEI pDNA, was freeze dried. The PEG PEI pDNA complicated with no FSH peptide was prepared by the similar technique. The encapsulating efficiencies in the complexes had been determined by gel retardation assay. The morphologies of your complexes were examined using the Joel Jem 2100 F transmission electron microscope, Particle size and zeta potential had been established applying the Malvern Zetasizer autosize 4700, Immunocytochemistry To detect the expression of FSHR and gro, immuno cytochemistry was employed.
Immediately after fixation with 4% parafor maldehyde, cells have been incubated with FSHR antibody or gro antibody overnight, and then in cubated with peroxidase conjugated anti rabbit IgG for 30 min. The staining reaction was carried out with di aminobenzidine. The cells had been counter stained with hematoxylin to detect nuclei, and imaged by light mi croscopy, Reverse selleck Ridaforolimus transcription polymerase chain response RNA was isolated from cells, and one ug of total RNA was reverse transcribed applying a cDNA synthesis kit according For actual time quantitative RT PCR, gro mRNA amounts were determined with SYBR Green detec tion in a two phase reaction employing the Eppendorf Mastercy cler ep Realplex RT PCR technique as described previously, Relative expression levels had been calculated employing the 2CT method and had been normalized to untreated control.
Enzyme linked immunosorbent Camostat Mesilate assay To investigate the gro ranges secreted by cells, cell su pernatants have been collected right after treatment method and examined using a gro ELISA kit according towards the manufacturers directions. Briefly, samples and standards had been additional to ELISA microplates and incubated for 1. five h at area temperature. Following washing, samples had been incubated with gro conjugate for 1 h at four C, and then with substrate so lution for 15 min at area temperature. The response was stopped employing quit answer. The optical density of ultimate solutions was measured employing a microplate reader at 450 nm wavelength. Cell proliferation analysis Cells were seeded into 96 well plates at a density of 1 ? 104 cells per nicely and incubated overnight. Cells have been in cubated with serum free medium containing 1. five ug of gro siRNA NP or FSH33 gro siRNA NP for 4 h. Medium was then replaced with fresh medium containing 10% fetal bovine serum. At 24 h, 48 h, 72 h and 96 h, 10 ul of CCK 8 solution was added and the cells incubated for one h.
This CK2 unique motif might let for the phosphorylation of substr
This CK2 precise motif may allow for your phosphorylation of substrates with proline or nonproline residues on the P 1 place from the sub strate resulting from its increased side chain flexibility, giving H bonding to the key chain oxygen with the strained place the CMGC arginine does, but in addition accom modating alterative H bonding, The conserved CK2 glycine following it might more contribute on the flex ibility allowing a higher array of most important chain conforma tions, whilst cgd7 1320 won’t conserve this glycine. The PfCK2a is characterized with pro tein kinase action. and gene disruption experiments have shown that it is critical on the asexual blood stage in Plasmodium, thereby delivering proof that it may be a drug target, Moreover, it’s differential susceptibility to a minor molecular inhibitor generating it an desirable target for antimalarial intervention, and poten tially its orthologues in other parasitic disorders.
Lastly, 3 uncharacterized CMGC members without identified orthologues outdoors of Cryptosporidium spp. had been also recognized. cgd1 810, selleck chemicals cgd7 4850, and cgd2 3890. Atypical group Atypical protein kinases lack sequence similarity to the eukaryotic protein kinase domain hidden Markov model profile and as this kind of are unrelated by sequence to ePKs. yet, they’ve got been shown experimentally to have protein kinase activ ity or are clear homologues of aPKs with demonstrated protein kinase action. One can find four C. parvum kinases which are identified as atypical primarily based on their P. falciparum and T. gondii orthologues, together with 2 RIOs and 2 ABC1s, There might be as several as 24 from T.
gondii and four from P. falciparum, TKL group In C. parvum, there are actually 3 TKL enzymes which includes cgd8 2430, cgd3 2900, and cgd3 4310 with just about every possessing P. falciparum and or T. gondii TKL orthologues, even though T. gondii and P. falciparum kinase inhibitor SCH 900776 every single includes 5 TKL enzymes, Lately, function on PfTKL 3 has demonstrated that it’s necessary for asexual parasite proliferation in human ery throcytes, On top of that, the authors showed that it undergoes in vitro autophosphorylation and phosphory lates exogenous substrates the two of that are dependent over the presence of the sterile a motif domain at the N terminus. Even though these C. parvum and P. falci parum TKL 3 orthologues only share 30% overall sequence identity, they each possess a SAM domain, at the same time as putative MORN motifs which can be N terminal to the kinase domain and not shared from the TgTKL 3 ortholo gue.
In the case in the P. falciparum orthologue, there may be an additional N terminal domain of 1300 residues upstream of your SAM domain, correspondingly this domain is only 300 residues from the C. parvum enzyme. Even though CpTKL 1 and PfTKL one also bear these SAM and MORN motifs, the third CpTKL doesn’t, OPK group You will discover 2 clades of protein kinases totally different to Apicomplexa, namely rhoptry kinases and FIKK kinases, We performed BLAST examination from the sequences of all known Toxoplasma ROP kinases against the C.
In dinoflagellate gene transcripts propose, by analogy with trypa
In dinoflagellate gene transcripts propose, by analogy with trypanosomes, the publish transcriptional manage of gene expression. Consistent with this particular hypothesis, several physiological processes are found to become regulated submit transcriptionally, Similarly, microarray ana lysis of acute tension responses in K. brevis didn’t reveal the activation of stress genes beneath situations exactly where they were proven to become induced with the pro tein degree, Microarray analyses noticed only 3% of contrast, the options unique to P addition that were signifi cantly enriched had been more members of GO classes corresponding to the ribosome or rRNA binding, the chlor oplast and photosynthesis, Early Responding Transcripts Possess the Spliced Leader Given the presence of the spliced leader sequence on all dinoflagellate nuclear encoded transcripts investigated to date, and its implication of submit transcriptional management of gene expression, it was relatively surprising to see the early response of gene transcripts in Cluster 5 of the two the N and P addition experiments, which had been domi nated by PPR proteins.
A few of these transcripts improved over 3 fold by 1 h following P addi tion, This led us to question regardless of whether these early reply ing transcripts represent a class of genes not processed via the SL mechanism and beneath transcriptional selleck chemical handle. Because the sequences from which probes about the array have been developed are ESTs or contigs representing incomplete gene sequences, we selected representative early responding PPR containing transcripts for confir mation of the 5 SL sequence by carrying out PCR implementing a SL primer and gene distinct primers.
Because of the substantial sequence similarity concerning PPR contigs in our K. brevis EST library, the reverse primers Asaraldehyde created towards contigs 183, 3257, 3556, and 3574, when paired using the SL pri mer, actually amplified many products. Thus, by way of cloning and sequencing we’ve recognized the presence in the SL on just more than forty contigs annotated as PPR pro teins. These benefits recommend the early responding genes are certainly not exceptions towards the SL trans splicing mechanism prevalent in dinoflagellates. Discussion The absence of recognizable promoter sequences on dinoflagellate genes along with the presence of 5 SL on transcripts transforming in excess of a circadian day in Pyrocystis lunula and K. brevis, Yet in K. brevis 10% of transcripts had been differentially expressed above a light dark cycle, a percentage that does not vary substantially from other photosynthetic eukaryotes that use common transcriptional controls, Since microarrays report only changes in transcript abundance, it stays unre solved by what mechanism these changes are achieved.
While genes while in the Wnt pathway appear to become usually dow
Though genes from the Wnt pathway appear to be typically downregulated with the postlarval stage, there is certainly no stage specific enrichment or depletion observed for your Notch and TGF B pathways. Transcription things that emerged in the metazoan lineage are believed to possess had a crucial role in growth and differentiation of cell styles, Differential expression of transcription things and transcriptional regulators suggest a critical part in coord inating the transformations that arise during the pela gobenthic transition.
About 86% of genes bearing regarded transcription element domains represented during the sponge genome have been also detected during the sponge transcriptome, Members of your bZIP and Tbox families are enriched in competent larvae find more information and may very well be involved in regulating the expression of genes necessary for settlement, However, in spite of upregulation of transcription aspect expression, the total mRNA transcripts in competent larvae didn’t change drastically from the precompetent stage, as evidenced through the properly correlated transcription profiles on the two stages, This observation suggests that other regulatory proteins current from the pelagic larvae are professional viding an extra layer of management above gene expres sion. One example is, genes encoding proteins similar to the repressors Ncor, Sin3, Tbl1xr, Phf, and Nacc, show a trend towards downregulation through the transition from larva to adult. Chromatin modifiers also show differen tial expression in pelagic versus benthic phases.
While histone acetylation genes are located at reduced ranges and histone deacetylases are really expressed in competent larvae, the opposite is real after settlement, Hence, competent sponge larvae seem for being poised for quick and widespread transcription on settlement, great post to read with several mechanisms in spot to control adjustments in gene expression. From the grownup sponge, which displays a drastically different transcription pro file from larvae, numerous transcription aspect groups, in cluding the zinc finger, forkhead, ETS, and homeobox domain containing elements, as well as other transcrip tional activators, are upregulated, Contributing to regulation of gene expression during the benthic stages are DNA methylation and histone acetylation proteins, at the same time as compo nents within the minor RNA machinery and RNA bind ing proteins which are upregulated upon settlement.
Genes which are selectively expressed in the very same devel opmental stage may perhaps be regulated through the identical set of transcription factors. For example,  brief chain collagens and members of the cadherin domain and scavenger re ceptor domain containing gene families, display coordinated upregulation on the adult stage, Within the adult, transcrip tionally lively genomic loci with three or extra scaven ger receptor domain containing genes are organized in tandem arrays and therefore are potentially regulated as a result of a shared cis regulatory element, Photosensory strategy Upon emerging from your brood chamber A.
brief chain collagens and members of the cadherin domain and scavenger re ceptor domain containing gene families, display coordinated upregulation on the adult stage, Within the adult, transcrip tionally lively genomic loci with three or extra scaven ger receptor domain containing genes are organized in tandem arrays and therefore are potentially regulated as a result of a shared cis regulatory element, Photosensory strategy Upon emerging from your brood chamber A.
Changes during the activ ity of various cell wall linked genes
Changes while in the activ ity of numerous cell wall associated genes were identified to result in the abnormal growth of juice sac granulation, whilst modifications in cell wall framework or during the parts of the membranes of the segments and juice sacs in the course of fruit growth and ripening plainly influenced the formation within the fruit pulp melting charac teristic, Among the cell wall related genes uncovered right here encoded a pectinesterase, an enzyme which modifies the assembly and disassembly of pectin, a frequent com ponent with the primary cell wall. In tomato, the gene for pectinesterase was tremendously expressed just before ripening, and was down regulated by ethylene as ripening starts, Right here, the expression on the gene encoding the pecti nesterase PPE8B precursor decreased because the WT fruit matured, when within the blood orange, the expression of a pectinesterase gene continues to be proven to increase during fruit improvement and ripening, It may very well be because of the distinct members recognized within the two studies.
The hemicellulose xyloglucan is usually a frequent element of your cell wall, and is hydrolysed and trans glycosylated kinase inhibitor Sunitinib by xyloglucan endotransglycosylase in develop ing tissues and ripening fruits, Right here, a gene encoding this enzyme was up regulated while in fruit improvement and ripening, indicating its probable part in cell wall degradation in the course of ripening. Both the early enlargement with the citrus fruit driven by cell expansion and the later on ripening procedure demand the presence of expansins to loosen the cell walls, and a number of genes encoding expansins had been detected during the current examine and their expression was greater with the later on stages of fruit advancement and ripening.
The accumulation of carbohydrates represents one of the most evident improvements which occur in the course of citrus fruit advancement and ripening. The perceived changes in expression of genes concerned in carbohydrate metabolism Sorafenib here had been steady together with the findings of other transcrip tomic analyses in citrus, The kind of sugar deposited to a high level within the cell vacuole in citrus is predomi nantly sucrose, unlike in grape, the place it is actually glucose and fructose, Reflecting this variation, the expression profile of your gene encoding sucrose synthase in citrus was rather numerous from that inside the grape, Nonetheless, in each species, sugar is essential for your regulation of colour improvement, and probably also for other ripening processes, It had been notable the gene encoding a sucrose phosphate synthase was induced all through fruit development and ripening, possibly since the activity of this enzyme can be expected for the re synthesis of sucrose to permit its even more transport to the vacuole, Of some curiosity also was the behaviour of the gene encoding valencene synthase, an enzyme the activ ity of which was regarded to become induced by ethylene, and which was element within the natural ripening method in citrus, Valencene is surely an essential part in the aroma with the ripe sweet orange fruit.

enysii and P fastigiatum tag library, respectively The library
enysii and P. fastigiatum tag library, respectively. The library sizes for each of your 6 lanes resulted from summing all locus counts for a certain lane and had been distinctive for every from the 10 datasets, To be able to investigate the influence of a the usage of a rela tively distant reference dataset and b the usage of partial contigs to the differential expression analysis we com pared the amount and overlap in the differentially expressed genes located with each and every of the ten information sets summarized in Additional file 3. Comparison of tag profiling datasets. Numbers of DEGs Making use of only full length transcripts of P. fastigiatum and making it possible for for no mismatch we inferred one,039 and one,239 differentially expressed genes for P. fastigiatum and P. enysii, respectively. When one particular mismatch was allowed from the P.
enysii tags these numbers enhanced to one,086 and 1,366, When mapping the tags with no more hints mismatches towards all out there P. fasti giatum contigs that have been longer than 200 bp, representing the leaf transcriptome of this species, 2,722 and two,702 genes were inferred for being differentially expressed, Interestingly, permitting for one mismatch in P. enysii led to a lessen during the quantity of DEGs in P. fastigiatum and a rise in P. eny sii, Applying the small Arabidopsis reference dataset, only really number of differentially expressed genes were recognized. 249, 532, and 684 DEGs had been inferred in P. fastigiatum and 395, 755, and 805 DEGs were inferred in P. enysii inside the A0, A1, and A2 datasets, respectively, When the tags have been mapped towards the 33,602 cDNA sequences, representing the finish A.
thaliana tran scriptome, these numbers enhanced to one,009, 1,978, and three,364 in P. fastigiatum and one,219, two,335, and 3,309 in P. enysii, Therefore in each the small along with the huge A. thaliana reference sets, the number of DEGs inferred improved with an raising number of mismatches permitted, This improve was stron ger inside the significant A. thaliana reference sets, Comparison of tag profiling datasets. selleckchem Agreements and discrepancies involving sets of DEGs When comparing DEGs inferred for distinct datasets it is crucial to not merely assess their quantity but also whether or not exactly the same genes are inferred to become up regulated among unique datasets. Such as, although the amount of DEGs inferred with datasets P0 and P1 sug gests a substantial degree of similarity, only 926 and 774 genes are up regulated in the two datasets for P. enysii and P. fas tigiatum, respectively. We computed the number of overlapping genes in pairwise comparisons of all ten datasets, For your genes up regulated in P. fas tigiatum, the highest overlap to the P0 dataset was 80% with PL0.
96% of those unigene sequences matched to model or ganisms Clust
96% of these unigene sequences matched to model or ganisms. Clusters of orthologous groups classification, Gene ontology and KEGG Overall, 9,920 sequences from 32,445 Nr hits, had a COG classification. Among the 25 COG categories, two,693 genes fell into the cluster for basic function prediction only. 1,316 gnes fell to the COG transcription group. one,305 genes had been categorized as owning a purpose within the posttranslational modification, protein turnover and chaperones COG group. one,250 of genes fell into the replication, recombination and repair COG group. Cell motility, extracellular structures and nuclear struc ture COG groups contained the fewest genes. We obtained Gene Ontology functional annota tion according for the Nr annotation. Based mostly on sequence homology, 13,317 sequences were categorized into 44 functional groups.
We identified in every single on the three principal classes of the GO classification that metabolic approach, cell element cell and catalytic selleck chemical PTC124 activity are dominant functions, respectively, we also no ticed a substantial percentage of genes through the classes binding, organelle and cellular system, and only a few genes through the functions of locomotion, cell killing, vir ion and virion portion. In complete, we assigned 14,462 sequences to 119 KEGG pathways as shown in Table two. The pathways most represented from the one of a kind sequences have been metabolic pathways, biosynthesis of secondary metabolites, spliceosome, phenylpropanoid biosynthesis, starch and sucrose metabolism and fla vonoid biosynthesis. These annotations deliver a useful resource for investigating distinct processes, functions and pathways in L.
gmelinii Imatinib genes. We think that genes within the KEGG classes meta bolic pathways and starch and sucrose metabolic process play a substantial purpose in plant growth and improvement. Path strategies such as flavonoid and phenylpropanoid biosynthesis are significant in plant pressure resistance. Phenylpropanoids are a group of secondary plant metabolites derived from phenylalanine and function as structural and signaling molecules. Phenylalanine is very first converted to cinnamic acid by deamination, which is followed by hydroxylation and many methylation ways to create coumaric acid together with other acids having a phenylpropane unit. Re duction with the CoA activated carboxyl groups of these acids lead to the synthesis of corresponding aldehydes and alcohols.
The alcohols are known as monolignols,  and are commencing components to the biosynthesis of lignin. These straightforward phenolic compounds are essential in plant defense towards fungi and herbivorous insects. Being a result, phenylpropanoid metabolic pathways perform an essential part in plant development, growth and defense responses against pathogen and herbivore attacks. Protein Coding Region Prediction In total, 32,047 and 2,771 unigenes had been predicted by BLASTx and ESTScan, respectively.
and are commencing components to the biosynthesis of lignin. These straightforward phenolic compounds are essential in plant defense towards fungi and herbivorous insects. Being a result, phenylpropanoid metabolic pathways perform an essential part in plant development, growth and defense responses against pathogen and herbivore attacks. Protein Coding Region Prediction In total, 32,047 and 2,771 unigenes had been predicted by BLASTx and ESTScan, respectively.
